
- 代表機関:自然科学研究機構 基礎生物学研究所
- 課題管理者:東島 眞一
- FAX:0564-55-7580
- 分担機関①:宇都宮大学 バイオサイエンス教育研究センター
- 分担機関②:国立遺伝学研究所 ゲノム・進化研究系 分子生命史研究室
- 分担機関③:長浜バイオ大学 バイオサイエンス学部
- 分担機関④(バックアップ):宮崎大学 地域資源創成学部
概要Overview
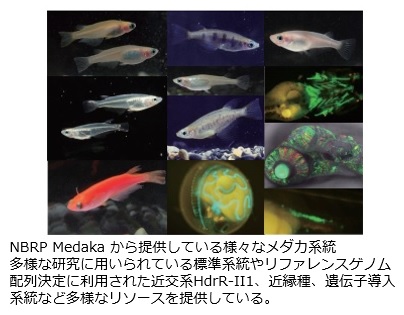 メダカは、世代時間が約3か月と短く、小型で飼育も容易です。またゲノムサイズは約800Mbpで、高精度な全ゲノム配列やゲノムアノテーションも整備されています。メダカは生息温度域や生息塩分濃度域が広いという生理学的な特性もあります。さらに、東南アジアを中心に様々な環境に適応した近縁種も利用できます。初期胚の操作に必須な孵化酵素も提供しています。
メダカは、世代時間が約3か月と短く、小型で飼育も容易です。またゲノムサイズは約800Mbpで、高精度な全ゲノム配列やゲノムアノテーションも整備されています。メダカは生息温度域や生息塩分濃度域が広いという生理学的な特性もあります。さらに、東南アジアを中心に様々な環境に適応した近縁種も利用できます。初期胚の操作に必須な孵化酵素も提供しています。
リソースの系統
・標準系統及び日本産、中国産、韓国産野生由来系統:約100系統
・突然変異系統及び遺伝子導入系統:約730系統
・近縁種:約10系統
・ゲノムリソース(cDNA, BAC, Fosmidクローン):約73万クローン
・孵化酵素
リソースの特徴
NBRP Medakaは世界規模で展開する唯一のメダカ生物遺伝資源センターです。生体リソース、ゲノムリソース、孵化酵素に加え様々なリソースのデータベースを提供しています。
代表機関での取り組み
基礎生物学研究所と宇都宮大学は東北大学と協力して野生由来系統のゲノム塩基配列を決定をします。またPacBio Hi-Fi readsを用いてより精度の高いリファレンスゲノムを構築します。
分担機関での取り組み
メダカ生命情報及びオミクスデータの統合に向けて、国立遺伝学研究所を中心に、利便性の高いウェブツールの開発に取り組んでいます
リソース関連プログラム課題
【ゲノム情報等整備/ゲノム情報等整備プログラム】
| 2023年度-2024年度 | 表現型可塑性を探るメダカゲノム基盤の整備 |
| 2022年度-2023年度 | メダカ野生由来系統のゲノム多型情報整備 |
| 2010年度 | メダカ近交系シークエンスによるゲノム多型情報の整備 |
| 2009年度 | メダカ完全長cDNAリソースの整備(両端読み234085件、全長配列決定2685件) |
| 2008年度 | 完全長cDNAリソースの整備 (8693件) |
| 2007年度 | 完全長cDNAリソースの整備 (全長読み5906クローン, 両端読み 265859件) |
| 2002年度 | ゲノム情報 – 成果:ゲノム情報等整備プログラム:2002年度-2006年度 |
【基盤技術整備/基盤技術整備プログラム】
| 2024年度-2025年度 | 次世代型メダカバイオリソース整備とその拠点形成 |
| 2012年度-2013年度 | 生殖細胞の凍結保存と借り腹生産による系統の回復に関する技術開発 |
| 2007年度-2009年度 | メダカ遺伝子機能解析汎用系統の開発 |
